Cool that Google embedded the entire PubMed dataset in BigQuery for semantic search, but....
Watch out for the sample queries on the site. It could cost you.
DISCLAIMER: I am relatively new to GCP and Bigquery, so please double check and offer clarifications. That helps us learn.
Google’s recent announcement about putting the entire PubMed database into BigQuery pre-embedded COMBINED with their recent announcement of MCP-ifying their services sounded too good to be true. Thirty-five million biomedical articles. Semantic search. The ability to create powerful Pubmed researcher agents on GCP. WITHOUT having to worry about the EXPENSIVE data engineering around maintaining my own embedding pipeline.
The second paragraph in Google’s announcement on Accelerate Medical Research in Bigquery describes this new service in BigQuery:
At Google Cloud, we’re addressing this challenge by making PubMed data available as a BigQuery public dataset with vector search capabilities from Vertex AI (both BigQuery and Vertex AI Vector Search are FedRAMP High authorized), enabling semantic search of medical concepts beyond simple keyword matching.
It likely costs thousands of dollars in compute time and significant overhead to keep a vector column up to date on 35 ‘million-ish’ articles to facilitate semantic search.
But Google has already done it. You don’t have to front the cost to vectorize the library (and maintain it).
So I asked myself….”well if BigQuery can now be an MCP server, then can I plug it into Claude Code, OpenWebUI, TypingMind, an OpenAI GPT, or wherever”?
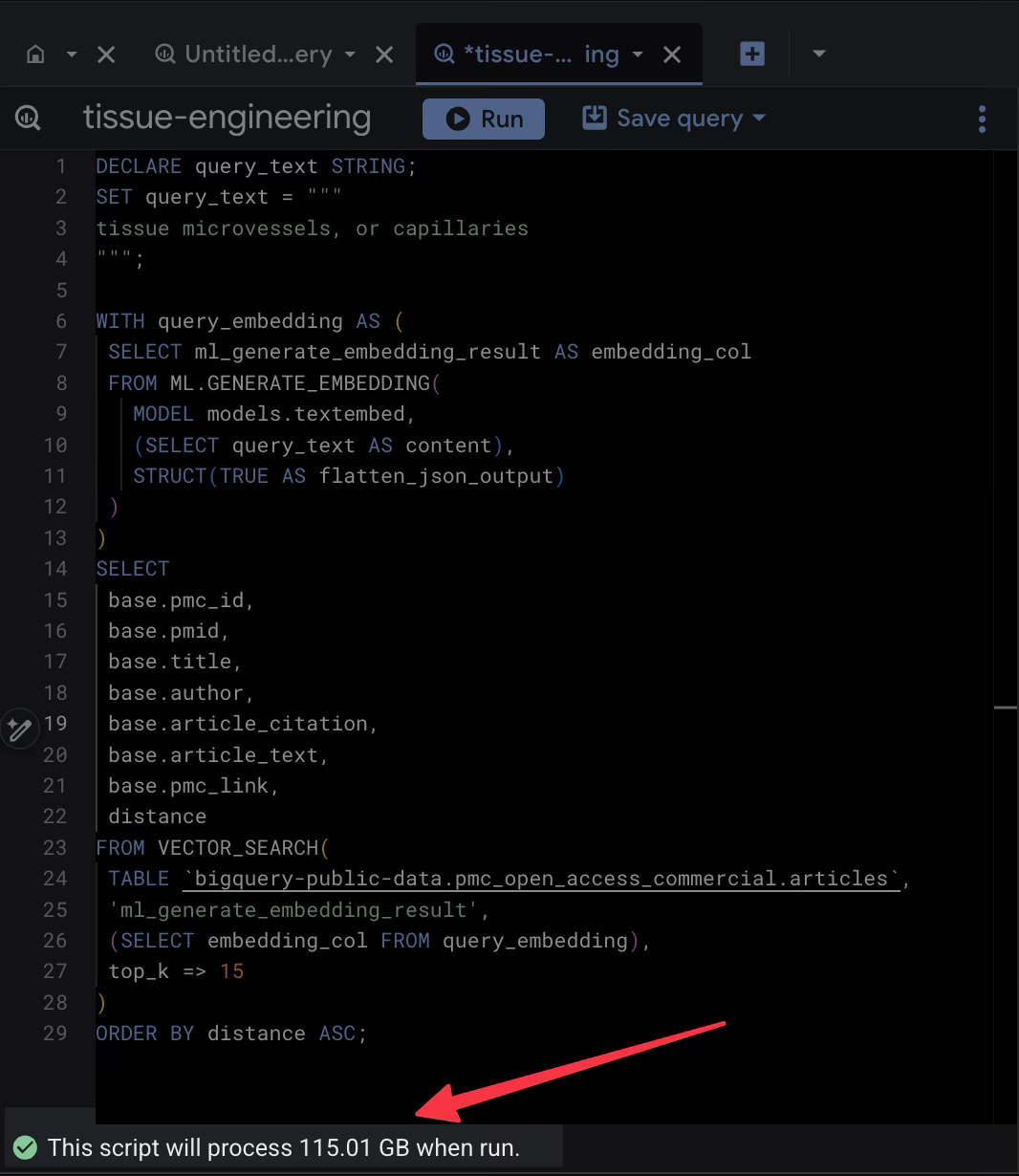
So off I went on this adventure fools errand. In my GCP account, I followed their "Getting Started" guide, copied their sample query, modified it to include research topics from one of my researchers and watched the magic happen.
Cool. It worked. BUT 115 GB SCANNED FOR ONE QUERY. That sounds expensive. The excitement waned.
For a single question. At BigQuery’s on-demand rate, that’s around 70c per query after free-tier. Run that a thousand times for a serious research project, and it wouldn’t take much to watch those precious research credits go up in smoke. Stacking that on top of agent costs likely won’t make leadership happy and optimistic about the value of AI.
What I found after research is that the difference between a pocket-emptying query and a practically free one came down to deleting a single line of code.
The Trap of the "Easy" Query
The problem lies in how BigQuery bills you. As it is mostly concerned about how data hungry you are, and the PubMed dataset is massive, the real weight is in the article_text column.
In the example Google provides, the SQL asks the database to perform a vector search to identify the most semantically similar articles to your query. But it also asks the database to return the full text of those articles in the same swoop.
This sounds efficient, right? It is not.
When I ran that standard query, BigQuery had to scan the embedding column to do the math, but because I requested SELECT base.article_text, it also decided to haul back that massive text column along for the ride. That’s what led to the 100+ GB of data.
The ~90% Discount
But if the vector search only needs the embeddings (the mathematical representation of the text) to figure out which articles match, does it really need the human-readable text until a human wants to dive deeper? Could that be the job of a second agent and some human in the loop process? Or a deterministic workflow?
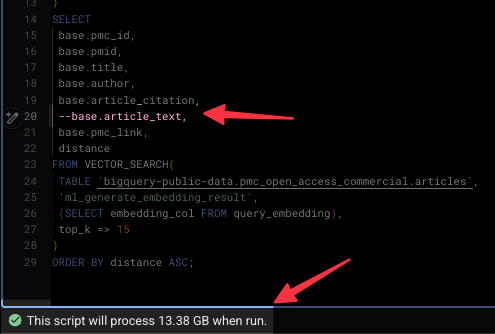
I started by the exact same query and deleted one line: base.article_text and hit run. The results came back just as fast, giving me the IDs, titles, and authors of the most relevant papers. But the data processed dropped from 115 GB down to 13ish GB.
By simply not pulling the full text through the computational pipeline, the price of the query dropped from a dollar to a dime. Those savings will add up.
Another Way to Build This
If you actually need the text (and spoiler: you probably do), maybe don’t ask for it during the search.
First, use the lean, 13GB query to find the top X Article IDs. That costs a few cents. Then, run a second, tiny query to fetch the full text for analysis 10 IDs. Cheaper.
It could probably even get better if you copy the table or just the rows you need to your own project (a one-time or maybe scheduled pull cost of a few dollars) and build your own vector index. Then you stop scanning GB and start scanning MB.
The Lesson
Cloud providers love to show you the "happy path" code that gets you a result in five seconds. FinOPS batteries are often not included. Watch out.
(Again: Please correct my misunderstandings of BigQuery and help me learn).
Ok now on to the MCP/A2A part of my journey….